- Research
- Open access
- Published:
CAZymes-based ranking of fungi (CBRF): an interactive web database for identifying fungi with extrinsic plant biomass degrading abilities
Bioresources and Bioprocessing volume 6, Article number: 51 (2019)
Abstract
Carbohydrate-active enzymes (CAZymes) are industrially important enzymes, which are involved in synthesis and breakdown of carbohydrates. CAZymes secreted by microorganisms especially fungi are widely used in industries. However, identifying an ideal fungal candidate is costly and time-consuming process. In this regard, we have developed a web-database “CAZymes Based Ranking of Fungi (CBRF)”, for sorting and selecting an ideal fungal candidate based on their genome-wide distribution of CAZymes. We have retrieved the complete annotated proteomic data of 443 published fungal genomes from JGI-MycoCosm web-repository, for the CBRF web-database construction. CBRF web-database was developed using open source computing programing languages such as MySQL, HTML, CSS, bootstrap, jQuery, JavaScript and Ajax frameworks. CBRF web-database sorts complete annotated list of fungi based on three selection functionalities: (a) to sort either by ascending (or) descending orders; (b) to sort the fungi based on a selected CAZy group and class; (c) to sort fungi based on their individual lignocellulolytic abilities. We have also developed a simple and basic webpage “S-CAZymes” using HTML, CSS and Java script languages. The global search functionality of S-CAZymes enables the users to understand and retrieve information about a specific carbohydrate-active enzyme and its current classification in the corresponding CAZy family. The S-CAZymes is a supporting web page which can be used in complementary with the CBRF web-database (knowing the classification of specific CAZyme in S-CAZyme and use this information further to sort fungi using CBRF web-database). The CBRF web-database and S-CAZymes webpage are hosted through Amazon® Web Services (AWS) available at http://13.58.192.177/RankEnzymes/about. We strongly believe that CBRF web-database simplifies the process of identifying a suitable fungus both in academics and industries. In future, we intend to update the CBRF web-database with the public release of new annotated fungal genomes.

Introduction
Annually, plants convert 100 billion tons of carbon dioxide and water to cellulose through photosynthesis (Field et al. 1998). Plant woody biomass is composed of carbohydrates (65–75%) (majorly separated into cellulose, hemicellulose and pectin), lignin (up to 20–30%), extraneous organic molecules (4–10%) and minerals (Pettersen 1984). Industries and researchers are continuously attempting new methods to explore and exploit plant woody biomass as a source of fermentable sugars (Carroll and Somerville 2009; Glass et al. 2013; Pauly and Keegstra 2008). However, the high recalcitrant nature and intricate networking properties of lignocellulose are major hurdles for industries to efficiently produce biofuels from plant biomass (Bischof et al. 2016; Himmel and Bayer 2009; Kubicek et al. 2009; Peterson and Nevalainen 2012). Previous studies have reported few bacterial strains and several wood-decaying fungi (classified as white rot, brown rot and soft rot fungi) have exhibited extrinsic lignocellulolytic abilities (Bischof et al. 2016; Peterson and Nevalainen 2012). Studies based on microbial degradation and conversion of plant biomass components to cellulosic ethanol and commercially important platform chemicals are rapidly gaining attention around the world (Abraha et al. 2019; Lijó et al. 2019; Lynd et al. 2008; Van Meerbeek et al. 2019; Zhu and Pan 2010). In the last decade several bacterial and fungal strains were identified and characterized around the world specifically to understand their lignocellulose degrading abilities (Bischof et al. 2016). Conventionally, the lignocellulolytic abilities of microorganisms are characterized in laboratories using various biochemical assays and methods. Although, by using the whole genome and transcriptome sequences we can completely understand the genome-wide distribution of CAZymes in a given fungi (Bischof et al. 2016; Lynd et al. 2008; Peterson and Nevalainen 2012; Zhu and Pan 2010).
Fungi secretes a variety of enzymes for the degradation of organic waste present on the earth’s surface thus, fungi play a crucial role in maintaining the nutritional cycles of our ecosystem (Glass et al. 2013). Compared to other microorganisms, filamentous fungi exhibit extrinsic plant biomass degrading abilities and industrially these fungi are highly important, e.g., food, beverage, pharmaceutical industries and also in production of commercially important enzymes (Himmel and Bayer 2009; Kubicek et al. 2009). Trichoderma reesei (Hypocrea jecorina), Aspergillus niger and Neurospora crassa are popular and commonly used industrial fungi for the production of commercial grade cellulases and several other highly catalytic enzymes. Recent fungal genomic studies strongly endorse the plant biomass degrading abilities of ascomycetes and basidiomycetous fungi (de Vries and Visser 2001; Dunlap et al. 2007; Glass et al. 2013). Development of public repositories such as JGI-MycoCosm, 1000 fungal genome project, Hungate genome projects have fueled the genome sequencing projects of several wood-decaying fungi and bacteria (Bischof et al. 2016; Lynd et al. 2008; Peterson and Nevalainen 2012; Zhu and Pan 2010). Understanding the genetic makeup of a microorganism in the first phase can reveal its plant biomass degrading abilities.
State of the art reviews have extensively reported about the plant cell wall-deconstructing enzymes in the past (Bateman and Basham 1976; de Vries and Visser 2001; Doi and Kosugi 2004; Glass et al. 2013; Minic and Jouanin 2006). Zhao et al. (2013) have compared the proteomic annotations of plant cell wall-degrading enzymes of 103 fungi representing different fungal phylum’s (Zhao et al. 2013). This study has reported about the genetic diversity of fungi, and the influence of fungal nutrition and host specificity on genome-wide composition of fungal CAZymes (Zhao et al. 2013). In our previous studies, we have compared the genome-wide distribution of CAZymes (carbohydrate-active enzymes) among the popular white rot, brown rot and soft rot fungi to understand and reveal their lignocellulolytic abilities and later ranked the fungi based on their degrading abilities. Similarly, we have also compared the genome-wide annotations of anaerobic fungi belonging to Neocallimastigomycota division to understand their plant cell wall-degrading and biohydrogen-producing abilities. Studies focused on understanding the plant cell wall-degrading abilities of individual fungal strains were reported in the last decade (Castillo et al. 2017; Geiser et al. 2016; Henske et al. 2017; Hüttner et al. 2017; Looi et al. 2017; Qin et al. 2017; Vidal-Melgosa et al. 2015; Zhang et al. 2016).
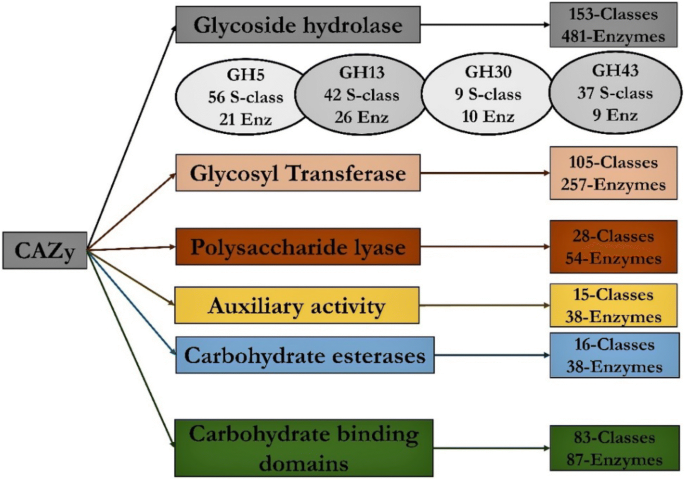
The microbial enzymes involved in plant cell wall-degradation are collectively called carbohydrate-active enzymes (CAZymes). These CAZymes were classified into six major groups as glycoside hydrolases, glycosyl transferases, polysaccharide lyases, carbohydrate esterases, auxiliary activity enzymes and carbohydrate-binding modules (Lombard et al. 2013). CAZy database has played a key role in enhancing present day’s knowledge about carbohydrate-active enzymes and it has influenced various groups around the world to pursue research on CAZymes. The primary understanding of CAZymes’ and their mode of action would significantly benefit the researchers identifying microorganisms with extrinsic plant cell wall-degrading enzymes (King et al. 2011; Kubicek et al. 2009; Zhao et al. 2013). However, the immenseness of the present day’s CAZy database makes it difficult for the researchers to specifically track down CAZymes based on the substrate they degrade. Biological contextualization of the whole genome sequencing studies of microorganisms using annotating databases such as InterPro (Hunter et al. 2008), dbCAN (Yin et al. 2012), CAT (Park et al. 2010) Hotpep (Halima 2019), provides the genome-wide CAZy annotations. However, the CAZyme annotations obtained are listed by their group and family numbers, e.g., GH1. Presently, to find all the enzymes that are classified in the GH1 family one must visit the CAZy web-database → select “Enzyme classes” → select “Glycoside Hydrolases” → select “GH1” and then specifically look for the enzyme of your interest.
In this study, we have reported a basic search page for finding and retrieving the complete list of CAZymes involved in biosynthesis and breakdown of carbohydrates. We have listed all the CAZymes groups and families that are involved in the degradation and conversion of plant cell wall components such as cellulose, hemicellulose, lignin, pectin, starch and inulin. This search page can be used as a primary material for understanding, exploring and designing CAZyme-based experiments. We have also reported a simple and efficient web-database for searching and short-listing the fungi based on CAZyme group or class which are involved in depolymerization of plant cell wall carbohydrates. To the best of our knowledge this study reports the first web-database with search functionalities for identifying a specific fungus based on its CAZyme coding abilities. Understanding the genome-wide distribution of CAZymes in first phase will significantly benefit the fungal laboratory and industrial-based projects.
Materials and methods
Design and construction of S-CAZymes
We have manually retrieved all the carbohydrate-active enzymes from the CAZy database (http://www.cazy.org/) which were originally classified under auxiliary activity, polysaccharide lyases, glucosyl transferases, glycoside hydrolases, carbohydrate esterases and carbohydrate-binding modules enzyme classes, respectively. The S-CAZy website contains three pages:
-
a.
Plant cell wall-degrading CAZymes (PCW-DE): as our primary research focus was to study about the plant cell wall component degrading CAZymes (such as cellulose, hemicellulose, lignin, pectin, chitin, starch and inulin degrading CAZymes).
-
b.
Search for CAZymes and CAZy families (S-CAZymes): we have provided a dynamic search functionality to search among all the listed CAZymes (full names) and their corresponding enzyme commission numbers, which enables us to retrieve the list of CAZymes involved in biosynthesis and depolymerization of carbohydrates.
-
c.
Enzyme activities (E.As) exhibited by carbohydrate-binding modules (CBM): similarly, in E.A-CBM webpage we have provided dynamic functionality to retrieve all the CBMs and their corresponding enzyme activities, E.A-CBM webpage lists the enzyme commission number, enzyme activities exhibited by CBM domains and all the corresponding CBM classes.
We have used HTML (hypertext markup language) version 5 majorly for developing our S-CAZy webpage. The webpage layout was designed and arranged using the bootstrap framework of CSS (cascading style sheets) https://maxcdn.bootstrapcdn.com/bootstrap/3.3.7/css/bootstrap.min.css. The dynamic search functionality for the S-CAZymes and E.A-CBM web pages were developed using the ajax google api https://ajax.googleapis.com/ajax/libs/jquery/3.3.1/jquery.min.js and bootstrap https://maxcdn.bootstrapcdn.com/bootstrap/3.3.7/js/bootstrap.min.js Java scripts, respectively. (http://13.58.192.177/RankEnzymes/SearchCAZymeCAZyClasses).
CBRF web-database design
As of today, the JGI-MycoCosm database resides 1057 fungal genome projects out of which 443 fungal genomes were published, publicly available and 613 fungal genomes are not published. In our present project, we have retrieved genome-wide distribution of CAZy annotations (CAZy groups and classes) of 443 published fungal genomes. Out of these 443 fungal genomes, 56 fungal genomes do not include CAZy annotations as of today. The CBRF web-database was designed with two major functionalities (a) sorting functions: ascending or descending orders and (b) sorting fungi based on CAZymes. The “sorting fungi based on CAZymes” function lists all the CAZy enzyme classes and groups (e.g., AA → AA1), these sorting options can also be selected using the dropdown option (which automatically pops up once user starts typing). Using the existing literature and based on our previous studies we have separated and classified the CAZymes based on the substrate they act such as: (a) cellulose, (b) hemicellulose, (c) lignin, (d) pectin, (e) starch and (f) inulin. We have provided the sorting functions for each of plant cell wall components (i.e., cellulose-C, hemicellulose-H, lignin-L, pectin-P, inulin-In and starch-S, respectively), e.g., the listed fungi are sorted based on their total cellulolytic ability (calculated by adding all the genes encoding for cellulolytic CAZymes). We have also provided a specific “Search-box” function to search for a specific fungus using its scientific name for comparing and analyzing its plant cell wall-degrading abilities.
CBRF construction
The above retrieved genome-wide fungal CAZymes data were initially segregated based on the genome code, scientific name, status of the work based on its availability (i.e., published or not published) assembly length and total number of genes. Later, genome-wide CAZy annotations including CAZy groups and classes of all the published fungi retrieved from JGI-MycoCosm database was considered for CBRF construction. These datasheets were primarily imported into MySQL database (version 8.0 CE) and Eclipse IDE photon platform, by creating a data schema using Spring data JPA and by creating model classes. We have created a rest service to create database table by using the inbuilt functionality for pulling the data from excel sheets using Apache POI jars and inbuilt POI method and frameworks. We have created a functionality named “search classifiers” for ranking the selected fungal genomes based on specific search of CAZyme. Later we have created various other services for saving, fetching, deleting and providing the data in user friendly graphical user interface. These user interface webpages were created using HTML, CSS, bootstrap, jQuery, JavaScript and ajax, which will indirectly call the services and render the responses on the webpages. We have exclusively provided functionalities such as sorting data based on specific search term, using the data tables and pagination frameworks of jQuery by creating responsive tables (http://13.58.192.177/RankEnzymes/about).
Results and discussion
Initial understanding about distribution of CAZymes in a selected microorganism’s genome will significantly help in conducting further experiments which benefits several microbe-based industries by developing cost- and time-efficient processes. As mentioned above in data retrieval section CAZy database is widely distributed, each CAZy family further includes several enzymes that are grouped based on their structural features (Busk and Lange 2013; Davies and Henrissat 1995; Henrissat 1991; Henrissat and Bairoch 1993, 1996; Henrissat and Davies 1997). Previous studies have extensively reported that microorganisms with wide variety of CAZymes can exhibit higher plant biomass degrading abilities (Daly et al. 2017; López et al. 2018; Min et al. 2017; Miyauchi et al. 2017; O’Connell et al. 2012) (Fig. 1).
Plant cell wall-degrading CAZymes (PCW-DE)
Using the pre-existing literature and CAZy web-repository, we have tentatively grouped the CAZymes based on the substrate they act upon. The CAZy groups and classes encoding for endoβ-(1→4) glucanases (cellulase), exo β-(1→4) glucanases (cellodextrinases), β-glucosidases, cellulase β-(1→4) cellobiosidase, lytic polysaccharide monoxygenases, cellobiose dehydrogenase, GMC-oxidoreductase and cellulose-binding domains were grouped as cellulose depolymerizing enzymes (Fig. 2). CAZy groups and classes encoding for endo-β-1,4-xylanase, β-glucosidase, β-mannosidase, β-xylosidase, Glucan β-1,3-glucosidase, mannan endo-β-1,4-mannosidase, alpha-l-arabinofuranosidase, xyloglucan-specific endo-β-1,4-glucanase, glucuronoarabinoxylan-specific endo-β-1,4-xylanase, arabinoxylan-specific endo-β-1,4-xylanase, acetyl xylan esterase, LPMO, xylan-binding modules, mannan-binding modules, arabinoxylan-binding modules and xyloglucan-binding modules were listed as hemicellulose-depolymerizing enzymes (Fig. 2).
Illustration of CAZy groups and classes involved in degradation of (a) cellulose, (b) hemicellulose, (c) lignin, (d) pectin, (e) starch and (f) inulin. For e.g., in CBRF web-database: cellulose degrading ability function calculates all the cellulose degrading CAZymes present in a given selected fungi and sorts all the fungi based on the number of cellulolytic CAZymes
Similarly, CAZy groups and classes encoding for laccase, p-diphenol oxygen oxidoreductase, ferroxidase, laccase-like multicopper oxidase, lignin peroxidase, manganese peroxidase, versatile peroxidase, peroxidase, aryl-alcohol oxidase, alcohol oxidase, pyranose oxidase, cellobiose dehydrogenase, GMC-oxidoreductases, vanillyl alcohol oxidase, alcohol oxidase, glyoxal oxidase, galactose oxidase, 1,4-benzoquinone reductase, Iron reductase, pyrroloquinoline quinone-dependent oxidoreductase, feruloyl esterase, cinnamoyl esterase, 4-O-methyl-glucuronoyl methylesterase and aryl esterase, carboxyl esterase were listed as lignin depolymerizing enzyme (Fig. 2) (Kameshwar and Qin 2019). The pectin-depolymerizing enzymes contained CAZy groups and classes encoding for polygalacturonase, exo-polygalacturonase, exo-poly-galacturonosidase, rhamnogalacturonase, rhamnogalacturonan α-1,2-galacturonohydrolase, α-l-rhamnosidase, exo-polygalacturonase, rhamnogalacturonan α-l-rhamnohydrolase, α-l-arabinofuranosidase, exo-α-l-1,5-arabinanase, β-galactosidase, pectate lyase, exo-pectate lyase, pectin Lyase, rhamnogalacturonan endolyase, rhamnogalacturonan exolyase, oligogalacturonate lyase, pectin methylesterase, pectin acetylesterase, rhamnogalacturonan acetylesterase, pectin acetylesterase, acetylesterase, pectin binding modules, galactan binding modules, l-rhamnose binding modules, arabinogalactan binding modules, respectively (Fig. 2). Finally, the CAZy groups and classes encoding for α-amylases, LPMO, α-glucosidase, starch phosphorylase, starch binding modules and endo-inulinase, exo-inulinase, inulin lyase and inulin binding module were listed for the starch and inulin depolymerizing enzymes, respectively (Fig. 2) (Kameshwar and Qin 2019).
Search-CAZymes (S-CAZy) and E.A-CBM webpages
The webpage: S-CAZy and enzymatic activities exhibited by CBM domains contain an efficient search option for finding a specific CAZyme from the complete CAZy classification. For instance, searching the S-CAZy webpage with the term “endo-beta-1,4-glucanase” would result in GH families GH5, GH7 and GH48, respectively. The S-CAZy webpage can also be used to find all the CAZymes involved in biosynthesis and breakdown of a specific substrate. For instance, the searching the S-CAZy webpage with first three letters of the term “manno”, “arabino” or “fructo” will result in all the CAZymes involved with biosynthesis or degradation of mannans, arabinans and fructosans (inulin), respectively. During its evolution glycoside hydrolases have separated into catalytic domain and non-catalytic carbohydrate-binding modules (CBM) which bind to the solid polysaccharides (Boraston et al. 2004). Previously, these polysaccharide recognizing polypeptides were named as cellulose-binding modules (CBM) or binding domains (CBD), however, due to their binding ability to a variety of polysaccharides, they were renamed as carbohydrate-binding modules (Boraston 1999). As of today, CAZy database hosts 83 CBM families, out of which cellulose-binding modules are distributed in 25 CBM families. CBMs can be majorly classified into three types: A-type (CBMs bind to surfaces of crystalline polysaccharides, e.g., cellulose, chitin), B-type (CBMs bind to internal longer sugar chains with more than four monosaccharide units) and C-type (CBMs bind to the ends of shorter sugar chains with not more than 3 monosaccharide units) (Armenta et al. 2017; Boraston et al. 2004; Gilbert et al. 2013). The “Enzymatic activities exhibited by CBM domains” webpage can be used for finding the carbohydrate-binding modules associated with a specific enzyme module. For e.g., searching E.A-CBM webpage with “cellulase” or with its E.C. number-3.2.1.4 will list all the CBM families associated with the cellulase enzymes, respectively. However, this page does not list the enzymatic activities corresponding to enzyme commission numbers “1.-.-.-.”, “2.4.1.-”, “2.4.2.-”, “3.1.1.-”, “3.2.1.-” and “4.2.2.-” as they do not correspond to a specific enzyme type.
Genome-wide proteomic studies of bacteria and fungi using microarray and RNA-sequencing results in huge data containing gene expression profiles which are either up or down regulated at cellular level. The RNA sequencing of various basidiomycetous fungi especially Phanerochaete chrysosporium, Postia placenta and several bacterial species have been already reported in the literature. The CAZy database (dbCAN) is widely used for annotating the newly sequenced genomes however, these genome-wide annotations are represented by their respective CAZy group and class number, e.g., GH1, GH2, GH45, etc. These CAZy group and class number does not reveal any additional information about the CAZyme or its function. At the same time, it is tedious to individually retrieve all the CAZymes based on the substrate they degrade from the CAZy database in a single step. The S-CAZy website can be used as a primer for specifically retrieving (a) all the plant cell wall-degrading CAZymes based on substrates; (b) classification (CAZy group and class) of a specific CAZyme in the CAZy database and (c) the basic information of the CAZymes of interest (Additional file 1) [Note: S-CAZymes webpage is just a simple search page to support and simplify the search process of CAZymes. However, for further information about carbohydrate-active enzymes (CAZymes) please visit: http://www.cazy.org/CAZy database].
Putative plant cell wall-degrading abilities
Recent genomic and proteomic studies of various plant cell wall-degrading fungi were centrally focused on lignocellulolytic CAZymes their occurrence and expression (Castillo et al. 2017; Geiser et al. 2016; Vidal-Melgosa et al. 2015; Zhang et al. 2016; Zhao et al. 2013). Knowing about the lignocellulolytic abilities of a microorganism prior to its experimentation will significantly benefit both the laboratory and industrial-based projects. Thus, in CBRF web database we have specifically added the simple functionalities for sorting fungi based on its plant cell wall-degrading abilities. We have retrieved the tentative CAZy groups and classes involved in cellulose, hemicellulose, lignin, pectin, starch and inulin depolymerization from the PCW-DE webpage of S-CAZymes. Using this information, we have tentatively developed a function for calculating all the genes encoding for CAZy groups and classes involved in degradation of plant cell wall components among all the 443 annotated fungal genomes. CBRF web-database displays all the fungi based on the selected sorting order. Using the CBRF web-database one can easily identify a specific fungus exhibiting extrinsic plant biomass degrading abilities (cellulose, hemicellulose, pectin, lignin, starch and inulin degrading CAZymes) by using the above-mentioned sorting functionality (Fig. 2).
CBRF framework, design and layout
As of today, approximately 11,264 studies include the term “plant cell wall degradation” in PubMed database, respectively. Out of which, 244 studies were related to plant cell wall-degrading microorganisms, 268 studies were based on CAZymes, 2985 studies related to plant cell wall degradation by fungi and 15 studies were based on plant cell wall-degrading fungal CAZymes, respectively (Fig. 3a). The JGI-MycoCosm, public repository presently hosts 1087-fungal genomes out of which 443-fungal genomes are published and 644-fungal genomes are sequenced but not published as of today (Fig. 3b) (Grigoriev et al. 2011, 2013). Based on their classification these 443 publicly available fungal genomes can be separated into 134 Basidiomycota, 264 Ascomycota, 7 Glomeromycota, 2 Mortierellomycotina, 12 Mucoromycotina, 2 Entomophthoromycotina, 6 Kickxellomycotina, 1 Blastocladiomycota, 3 Chytridiomycota, 5 Neocallimastigomycota, 8 Microsporidia and 1 Cryptomycota (Fig. 3b). However, out of these 443 fungal genomes, 386 genomes possess CAZy annotations and 57 genomes do not possess CAZy annotations as of today (Fig. 3b). The CBRF web-database was developed in the eclipse IDE photon platform using the Spring data JPA, Apache POI and Maven frameworks. The CBRF web database can be specifically used for sorting and searching fungi either by using a specific CAZy group or class or by using a specific plant biomass degrading ability, e.g., cellulose, hemicellulose, pectin, lignin, starch and inulin degrading abilities (Fig. 3a). Thus, obtained list can be easily downloaded in either of the file formats excel (.xls or .csv), PDF and user can also print or copy the results from the CBRF web-database. The CBRF web-database displays additional necessary information including genome code, complete scientific name, cellulose (C), hemicellulose (H), lignin (Li), pectin (Pe), starch (St), inulin (In), search CAZy term (e.g., AA), assembly length (A. Len), gene, total enzymes (#Enz) and published (Pub), respectively (Figs. 4a, b, 5).
Snapshot of CBRF-CAZymes-based ranking of fungi web database showing different sorting functionalities for retrieving the fungal information based on the search of interest [1—sorting functionality (provides two sorting options—ascending order and descending order); 2—the final sorted list of fungi can be exported in different file formats including: in.CSV, M.S. Excel, .PDF or print; 3—“Search functionality” to quickly search for a specific fungus of user’s interest; 4—lists the scientific names of all the top-sorted fungi, e.g.: in this window it only lists top 10 as we have selected only to show top 10 fungi; 5—we have specifically included the total fungal ability to degrade cellulose (C); 6-hemicellulose (H), lignin (L), pectin (P), starch (S) and inulin (In)]
Conclusion
The CBRF web-database was developed to identify and compare a specific fungus based on its plant cell wall-degrading abilities. The searching, and sorting functions of CBRF web-database enables users to find fungi both locally and dynamically among the 443 published fungal genomes. We intend to update the CBRF web-database regularly with the release of new published fungal genome sequences in the JGI-MycoCosm database. We have also developed a simple search page for understanding and finding a specific CAZyme and its classification among different CAZy groups and families. The S-CAZymes search page can also be used for finding all the enzymes involved in the biosynthesis and breakdown of a carbohydrate substrate. We also have reported about all the CAZy groups and families which are tentatively involved in synthesis and breakdown of the plant cell wall components. S-CAZymes search page can be used as a primer or preliminary material for understanding the CAZymes distribution and CAZymes required for the degradation and biosynthesis of plant cell wall polysaccharides. Our future works include updating CBRF web-database with the recently published open access fungal genomes by retrieving their genome-wide CAZy annotations. Similarly, we also intend to update the S-CAZymes webpage based on the carbohydrate-active enzymes classification updates of CAZy website.
Availability of data and materials
The data used and described in this paper can be freely accessed by using the CBRF web site at http://13.58.192.177/RankEnzymes/about through the latest versions of Google Chrome, Mozilla Firefox, Microsoft Internet Explorer and Apple Safari. The CBRF source code developed can be provided upon request.
References
Abraha M, Gelfand I, Hamilton SK, Chen J, Robertson GP (2019) Carbon debt of field-scale Conservation Reserve Program grasslands converted to annual and perennial bioenergy crops. Environ Res Lett 14(2):024019
Armenta S, Moreno-Mendieta S, Sánchez-Cuapio Z, Sánchez S, Rodríguez-Sanoja R (2017) Advances in molecular engineering of carbohydrate-binding modules. Proteins Struct Funct Bioinform 85:1602–1617
Bateman D, Basham H (1976) Degradation of plant cell walls and membranes by microbial enzymes. In: Physiological plant pathology. Springer, Berlin, pp 316–355
Bischof RH, Ramoni J, Seiboth B (2016) Cellulases and beyond: the first 70 years of the enzyme producer Trichoderma reesei. Microb Cell Fact 15:106
Boraston A (1999) Carbohydrate-binding modules: diversity of structure and function. In: The 3rd carbohydrate bioengineering meeting. The Royal Society of Chemistry, Newcastle upon Tyne, 1999
Boraston AB, Bolam DN, Gilbert HJ, Davies GJ (2004) Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J 382:769–781
Busk PK, Lange L (2013) Classification correlating to function of carbohydrate-active enzymes by recognition of short, conserved peptide motifs. Appl Environ Microbiol. https://doi.org/10.1128/AEM.03803-12
Carroll A, Somerville C (2009) Cellulosic biofuels. Ann Rev Plant Biol 60:165–182
Castillo L, Plaza V, Larrondo LF, Canessa P (2017) Recent advances in the study of the plant pathogenic fungus Botrytis cinerea and its interaction with the environment. Curr Protein Pept Sci 18:976–989
Daly P et al (2017) Transcriptomic responses of mixed cultures of ascomycete fungi to lignocellulose using dual RNA-seq reveal inter-species antagonism and limited beneficial effects on CAZyme expression. Fungal Genet Biol 102:4–21
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Structure 3:853–859
de Vries RP, Visser J (2001) Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol Mol Biol Rev 65:497–522
Doi RH, Kosugi A (2004) Cellulosomes: plant-cell-wall-degrading enzyme complexes. Nat Rev Microbiol 2:541
Dunlap JC et al (2007) Enabling a community to dissect an organism: overview of the Neurospora functional genomics project. Adv Genet 57:49–96
Field CB, Behrenfeld MJ, Randerson JT, Falkowski P (1998) Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281:237–240
Geiser E, Reindl M, Blank LM, Feldbrügge M, Wierckx N, Schipper K (2016) Activating intrinsic CAZymes of the smut fungus Ustilago maydis for the degradation of plant cell wall components. Appl Environ Microbiol. https://doi.org/10.1128/AEM.00713-16
Gilbert HJ, Knox JP, Boraston AB (2013) Advances in understanding the molecular basis of plant cell wall polysaccharide recognition by carbohydrate-binding modules. Curr Opin Struct Biol 23:669–677
Glass NL, Schmoll M, Cate JH, Coradetti S (2013) Plant cell wall deconstruction by ascomycete fungi. Annu Rev Microbiol 67:477–498
Grigoriev IV et al (2011) Fueling the future with fungal genomics. Mycology 2:192–209
Grigoriev IV et al (2013) MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res 42:D699–D704
Halima NB (2019) Prediction and analysis of GH14 family β-amylases in oat seedling extract: structure and function insights using in silico approaches. Int J Biol Macromol 125:361–369
Henrissat B (1991) A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 280:309–316
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788
Henrissat B, Bairoch A (1996) Updating the sequence-based classification of glycosyl hydrolases. Biochem J 316:695
Henrissat B, Davies G (1997) Structural and sequence-based classification of glycoside hydrolases. Curr Opin Struct Biol 7:637–644
Henske JK et al (2017) Transcriptomic characterization of Caecomyces churrovis: a novel, non-rhizoid-forming lignocellulolytic anaerobic fungus. Biotechnol Biofuels 10:305
Himmel ME, Bayer EA (2009) Lignocellulose conversion to biofuels: current challenges, global perspectives. Curr Opin Biotechnol 20:316
Hunter S et al (2008) InterPro: the integrative protein signature database. Nucleic Acids Res 37:D211–D215
Hüttner S et al (2017) Combined genome and transcriptome sequencing to investigate the plant cell wall degrading enzyme system in the thermophilic fungus Malbranchea cinnamomea. Biotechnol Biofuels 10:265
Kameshwar AKS, Qin W (2019) Systematic review of publicly available non-Dikarya fungal proteomes for understanding their plant biomass-degrading and bioremediation potentials. Bioresour Bioprocess 6:30
King BC, Waxman KD, Nenni NV, Walker LP, Bergstrom GC, Gibson DM (2011) Arsenal of plant cell wall degrading enzymes reflects host preference among plant pathogenic fungi. Biotechnol Biofuels 4:4
Kubicek CP, Mikus M, Schuster A, Schmoll M, Seiboth B (2009) Metabolic engineering strategies for the improvement of cellulase production by Hypocrea jecorina. Biotechnol Biofuels 2:19
Lijó L, González-García S, Lovarelli D, Moreira MT, Feijoo G, Bacenetti J (2019) Life cycle assessment of renewable energy production from biomass. In: Life cycle assessment of energy systems and sustainable energy technologies. Springer, Berlin, pp 81–98
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2013) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495
Looi HK et al (2017) Genomic insight into pathogenicity of dematiaceous fungus Corynespora cassiicola. PeerJ 5:e2841
López SC, Peng M, Issak TY, Daly P, de Vries RP, Mäkelä MR (2018) Induction of plant cell wall degrading CAZyme encoding genes by lignocellulose-derived monosaccharides and cellobiose in the white-rot fungus Dichomitus squalens. Appl Environ Microbiol AEM.00403-00418
Lynd LR et al (2008) How biotech can transform biofuels. Nat Biotechnol 26:169
Min B, Park J-H, Park H, Shin H-D, Choi I-G (2017) Genome analysis of a zygomycete fungus Choanephora cucurbitarum elucidates necrotrophic features including bacterial genes related to plant colonization. Sci Rep 7:40432
Minic Z, Jouanin L (2006) Plant glycoside hydrolases involved in cell wall polysaccharide degradation. Plant Physiol Biochem 44:435–449
Miyauchi S, Navarro D, Grisel S, Chevret D, Berrin J-G, Rosso M-N (2017) The integrative omics of white-rot fungus Pycnoporus coccineus reveals co-regulated CAZymes for orchestrated lignocellulose breakdown. PLoS ONE 12:e0175528
O’Connell RJ et al (2012) Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat Genet 44:1060
Park BH, Karpinets TV, Syed MH, Leuze MR, Uberbacher EC (2010) CAZymes Analysis Toolkit (CAT): web service for searching and analyzing carbohydrate-active enzymes in a newly sequenced organism using CAZy database. Glycobiology 20:1574–1584
Pauly M, Keegstra K (2008) Cell-wall carbohydrates and their modification as a resource for biofuels. Plant J 54:559–568
Peterson R, Nevalainen H (2012) Trichoderma reesei RUT-C30—thirty years of strain improvement. Microbiology 158:58–68
Pettersen RC (1984) The chemical composition of wood. Chem Solid Wood 207:57–126
Qin S, Ji C, Li Y, Wang Z (2017) Comparative transcriptomic analysis of race 1 and race 4 of Fusarium oxysporum f. sp. cubense induced with different carbon sources. G3 Genes Genomes Genet. https://doi.org/10.1534/g3.117.042226
Van Meerbeek K, Muys B, Hermy M (2019) Lignocellulosic biomass for bioenergy beyond intensive cropland and forests. Renew Sustain Energy Rev 102:139–149
Vidal-Melgosa S et al (2015) A new versatile microarray-based method for high-throughput screening of carbohydrate-active enzymes. J Biol Chem. https://doi.org/10.1074/jbc.M114.630673
Yin Y, Mao X, Yang J, Chen X, Mao F, Xu Y (2012) dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40:W445–W451
Zhang S et al (2016) De novo analysis of Wolfiporia cocos transcriptome to reveal the differentially expressed carbohydrate-active enzymes (CAZymes) genes during the early stage of sclerotial growth. Front Microbiol 7:83
Zhao Z, Liu H, Wang C, Xu J-R (2013) Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genomics 14:274
Zhu J, Pan X (2010) Woody biomass pretreatment for cellulosic ethanol production: technology and energy consumption evaluation. Bioresour Technol 101:4992–5002
Acknowledgements
We sincerely thank Hari Krishna Manupadi, Sai Sudha Anupindi for their valuable suggestions.
Funding
This work was supported by Natural Sciences and Engineering Research Council of Canada Funding (RGPIN-2017-05366) to Wensheng Qin and Mitacs Globalink Research Scholarship award to Ayyappa Kumar Sista Kameshwar.
Author information
Authors and Affiliations
Contributions
AKSK is involved in developing, executing and writing the complete framework of CBRF web-database, S-CAZymes webpage. LPR and WQ are involved in designing, guiding through the analysis section, improving and revising the manuscript. AKSK, LPR and WQ are involved in writing and reviewing the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
No applicable.
Consent for publication
Authors agreed to publish this article.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1.
Additional information explains the layout of the CBRF web-database and detailed explanation of comparison of CAZyme search process.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kameshwar, A.K.S., Ramos, L.P. & Qin, W. CAZymes-based ranking of fungi (CBRF): an interactive web database for identifying fungi with extrinsic plant biomass degrading abilities. Bioresour. Bioprocess. 6, 51 (2019). https://doi.org/10.1186/s40643-019-0286-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40643-019-0286-0